R
By: Alison Hu, Ethan Lee & Kelsey Wu
Difference between Python and R
R is an open source programming language and software environment. Much like Python, R can be used to do basic coding and data analysis using various available packages. The difference comes when looking at each language's overall purpose: R was created and developed mostly for statistical analysis, with much of the language filled with statistics terminology, while Python is more of a general-purpose language. Both R and Python are powerful languages for data analysis and visualization, but small discrepancies between the two (like R beginning vector indices at 1) also make them quite different.
Installing RStudio + Setup
You can install RStudio, the integrated development environment for R, at https://rstudio.com/products/rstudio/download/#download
Layout of RStudio environment
Once you have RStudio fully installed, open it and create a new R Script document by going to "File -> New File -> R Script".
Your workspace should look something like this:
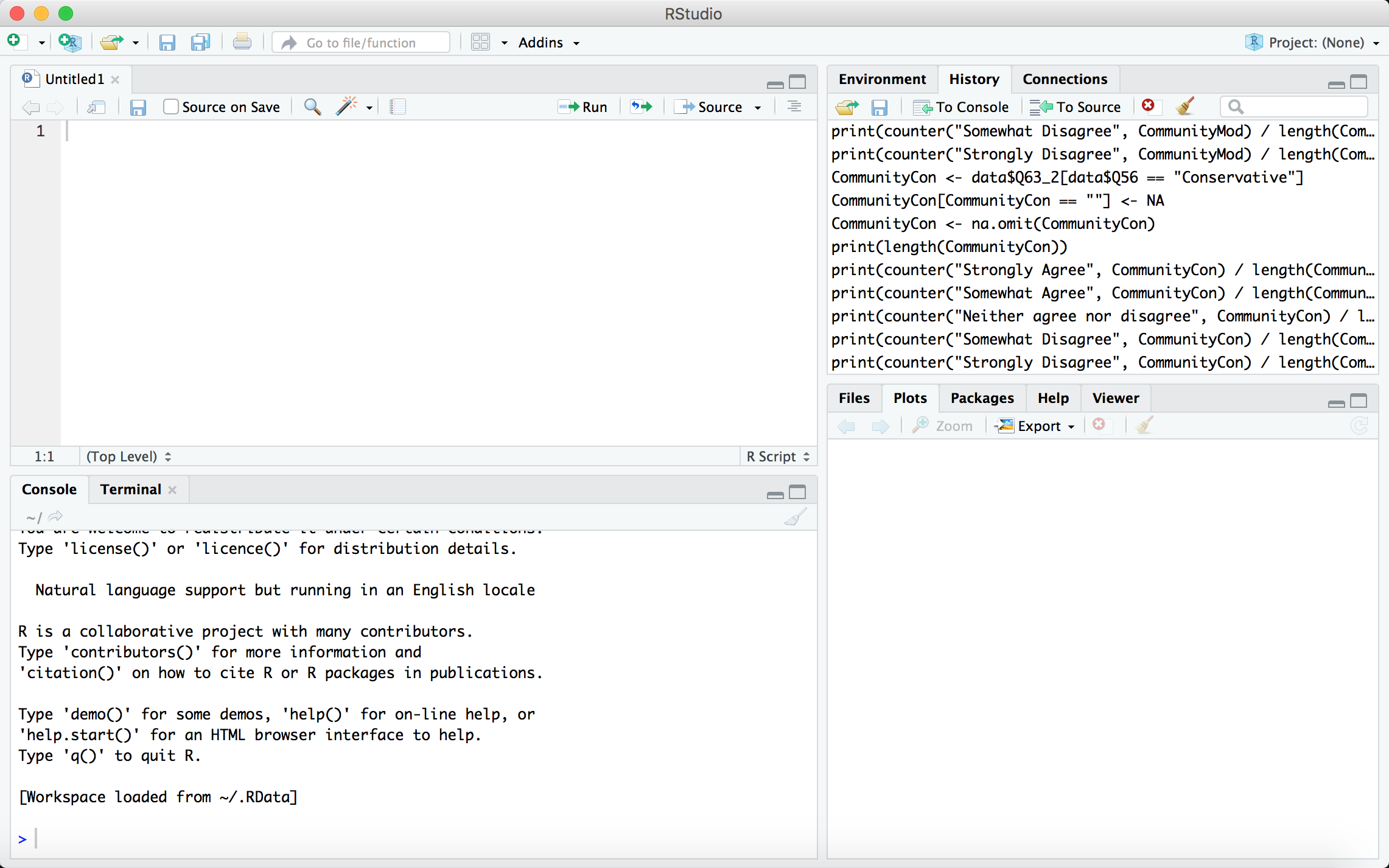
In RStudio, you're primarily working with four panels. At the top left is your code editor, where you can write out code in R and save it as a file (typically R Script or Markdown). At the top right, the "Environment" tab shows what variables/objects you have defined in your current environment, while the "History" tab shows the R commands you last conducted. At the bottom left, the "Console" tab is the R Console where you can write out R commands and see the results of all R code that is processed, while the "Terminal" provides access to a new terminal session from within the RStudio IDE. Lastly, at the bottom right, the "Files" tab shows files in your working directory, the "Plots" tab shows any plots you've created in your R session, and the "Packages" tab shows the R packages available to you.
Basics
Simple calculations
Similar to other programming languages, R can perform simple math calculations, both directly in the console and as a part of your script. The basic operators are: + (addition), - (subtraction), * (multiplication), / (division), ^ (exponentiation), and %% (modulo). For example,
outputs
There are also a number of built-in basic math functions, including abs(x) (absolute value of x), sqrt(x) (squart root), log10(x) (logarithm of x base 10), cos(x), sin(x), and tan(x). A full list can be found here.
Variables
Simply put, a variable stores a value in memory. In R, we use the '<-' symbol to assign a value to a variable. For example,
stores the number 2020 into a variable called "year."
Variable names must begin either with a letter or a period and can include any combination of letters, numbers, periods, and underscores; no other special characters or symbols may be used.
important
R is case-sensitive, so name (and access) your variables accordingly!
caution
Reserved words (if, else, while, break, function, etc.) have special uses in R and cannot be used as variable names.
Basic Data Types
Variables can have different types. In R, there are three basic data types: numeric, character, and logical.
Numeric
In the above example, "year" is of numeric type. Numeric types can store integers or floats (numbers involving decimals).
Character
Below, the variable "x" is of character type. Character types can be created with either single- or double-quotes.
Logical
Below, the variable "y" is of logical type. Logical types can store either TRUE or FALSE.
Data Structures
From these basic data types, we can start to build more complex structures in R.
Vectors
A vector contains elements all of the same data type. Vectors are created by wrapping elements inside c(). For example,
is a vector containing all character types.
Lists
A list contains elements of multiple data types. Lists are created by wrapping elements inside list(). For example,
contains numeric, character, and logical types.
Factors
A factor represents the categories present in a vector. Factors can be created by wrapping a vector inside factor(). For example,
creates the factor "my.factor". Calling levels(my.factor) would yield
Matrices
A matrix is a vector represented in a two-dimensional rectangular format. Matrices can be created by wrapping a vector inside matrix() and specifying the dimensions of the matrix, as follows:
This will create a two-by-two matrix containing the letters of "HODP".
Arrays
An array extends the idea of a matrix into multiple dimensions. Arrays can be created by wrapping a vector inside array() and specifying the dimensions of the array, as follows:
This will create a three two-by-two matrices containing the letters of "HODP".
Data Frames
A data frame is similar to a matrix, but each column can store a different data type. Data frames can be created by wrapping vectors inside data.frame(). For example,
creates a data frame with two columns, one labeled "letters" and one labeled "numbers", each with four rows.
Reading in files
Cleaning datasets (NA values)
To get a basic idea of how cleaning might work with a dataset in R, imagine we're working with the following vector v:
Perhaps v is a set of responses from a survey, and certain responses caused the non-numeric "not available" value NA to appear. If we simply try to the find the mean of v, we get the following result:
To get the mean of only numeric responses, we can use the "na.omit" function, which returns its input vector with NA values omitted.
Now, we can find the mean of just the numeric values!
Now, let's apply this idea of dropping NA values to an actual dataset.
Before you start cleaning your dataset, it's a good idea to get a feel for the data. The tidyverse package contains many very useful functions, including omitting NA values, filtering variables, sorting variables, etc.
To install the tidyverse package, type the following into the console:
To import a dataset into R, place the csv/Excel file in the same folder as your R file. Then, use the following command:
This means you are reading in a csv titled "telecom.csv", and you are assigning this dataset the reference "df". When you manipulate the dataset later, you will refer to this dataset as "df".
To take a quick look at your data, you can use the following functions:
Then, to omit NA values, you can use "drop_na" function, which drops rows that contain missing values.